| | Fang Bai Ph.D. |
Assistant Professor
Principal Investigator
Laboratory: Lab of Drug Design and Development Shanghai Institute for Advanced Immunochemical Studies
ShanghaiTech University; School of Life Science and Technology Email: baifang@@shanghaitech.edu.cn

Education
2004-2009: Dalian University of Technology, BE in Chemical Engineering and BA in English; 2009-2014: Dalian University of Technology(Jointly educated by Shanghai Institute of Materia Medica, Chinese Academy of Sciences)

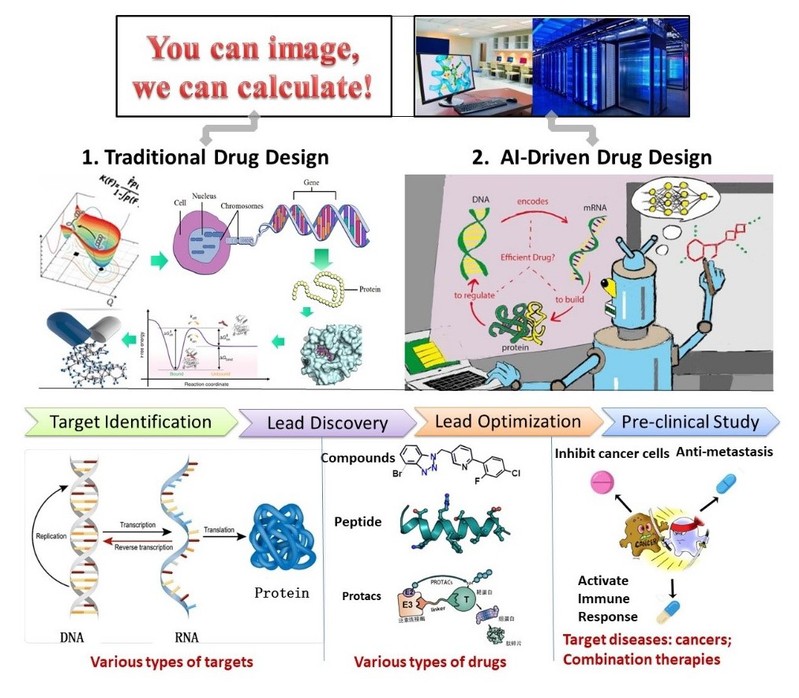
Employment 2014.8-2019.5 Postdoctoral Research Fellow, Rice University; 2019.5-2019.9 Assistant Professor of Research, University of Texas Health Science Center at Houston; 2019.10-present Assistant Professor, ShanghaiTech University  Research Interests 1. Developing computational drug discovery and design methodologies by combining artificial intelligence algorithms and theories of computational drug design methods, especially focusing on fragment-based drug design, PROTACs design and large-scale conformational sampling of proteins. 2. Cancer drug discovery and design, mainly targeting against tumor metastasis and immune escape.
 Selected Publications A. Amir1, F. Bai1,Y. Sohn, L. Song, S. Tamir, H. Marjault, G. Mayer, O. Karmi, P. Jennings, R. Mittlerf, J.N. Onuchic, A. Friedler, and R. Nechushtai. The anti-apoptotic proteins NAF-1 and iASPP interact to drive apoptosis in cancer cells. Chem. Sci. 2019, 10: 665-673(1Co-first Author). F. Bai, X. Pi, P. L, P. Zhou, H. Yang, X. Wang, M. Li, Z. Gao, H. Jiang. A Statistical Thermodynamic Model for Ligands Interacting With Ion Channels: Theoretical Model and Experimental Validation of the KCNQ2 Channel. Front. Pharmacol. 2018, 9:150. F. Bai, K. Liu, H. Li, J. Wang, J. Zhu, P. Hao, L. Zhu, S. Zhang, L. Shan, W. Ma, W. Zhang, H. Li, A.M. Bode &Z. Dong. Veratraimine modulates AP-1-depdendent gene transcription by directly binding to programmable DNA. Nucleic Acids Res. 2017, 46:546-557. F. Bai, F. Morcos, RR. Cheng, H. Jiang and J.N. Onuchic. Elucidating the druggable interface of protein-protein interactions using fragment-docking and coevolutionary analysis. Proc. Natl. Acad. Sci. USA. 2016, 113: E8051–E8058. F. Bai, F. Morcos, Y.S. Sohn, Darash-Yahana M., C.O. Rezende, C.H. Lipper, M.L. Paddock, L. Song, Y. Luo, S.H. Holt, S. Tamir, E.A. Theodorakis, P.A. Jennings, J.N. Onuchic, R. Mittler, and R. Nechushtai. The Fe-S cluster-containing NEET proteins mitoNEET and NAF-1 as chemotherapeutic targets in breast cancer. Proc. Natl. Acad. Sci. USA. 2015, 12: 3698-3703. F.Bai, S. Liao, J. Gu, H. Jiang, X. Wang, H. Li. An accurate metalloprotein-specific scoring function and molecular docking program devised by a dynamic sampling and iteration optimization strategy. J. Chem. Inf. Model. 2015, 55: 833-847. F.Bai, Y.Xu, J.Chen, Q.Liu, J.Gu, X.Wang, J.Ma, H. Li, J.N. Onuchic, and H.Jiang. Free energy landscape for the binding process of Huperzine A to acetylcholinesterase.Proc. Natl. Acad. Sci. USA. 2013, 110:4273-4278. F.Bai, H. Liu, L. Ton, W. Zhou, L. Liu, Z. Zhao, X. Liu, H. Jiang, X. Wang, H. Xie, and H. Li Discovery of novel selective inhibitors for EGFR-T790M/L858R. Bioorg. Med. Chem. Lett. 2012, 22:1365-1370. F.Bai, X. Liu, J. Li, H. Zhang, H. Jiang, X. Wang, and H. Li. Bioactive Conformational Generation of Small Molecules: A Comparative Analysis between Force-Field and Multiple Empirical Criteria Based Methods.BMC Bioinformatics 2010, 11:545. C. Lipper, J. Stofleth, F. Bai, Y. Sohn, S. Roy, R. Mittler, R. Nechushtai, J.N. Onuchic, P. Jennings. Redox-dependent gating of VDAC by mitoNEET. Proc. Natl. Acad. Sci. USA. 2019, 116: 19924-19929. Z. Tan, L. Xiao, M. Tang, F. Bai, J. Li, L. Li, F. Shi, N. Li, Y. Li, Q. Du, J. Lu, X. Weng, W. Yi, H. Zhang, J. Zhou, Q. Gao, J.N. Onuchic, A.M. Bode, X. Luo, Y. Cao. Targeting CPT1A-mediated fatty acid oxidation sensitizes nasopharyngeal carcinoma to radiation therapy. Theranostics 2018; 8:2329-2347. K. Yim, T. Prince, S. Qu, F. Bai, P. Jennings, J.N. Onuchic, E. Theodorakis and L. Neckers Gambogic acid identifies an isoform-specific druggable pocket in the middle domain of Hsp90β.Proc. Natl. Acad. Sci. USA. 2016, 113:E4801-E4809. M. Yahana, Y. Pozniak, M. Lu, Y. Sohn, O. Karmi, S. Tamir, F. Bai, L. Song, P. Jennings, E. Pikarsky, T. Geiger, J.N. Onuchic, R. Mittler and R. Nechushtai. Breast cancer tumorigenicity is dependent on high expression levels of NAF-1 and the lability of its Fe-S clusters. Proc. Natl. Acad. Sci. USA. 2016,113:10890-10895 Wang, Y. Jiang, J. Ma, H. Wu, D. Wacker, V. Katritch, G. Han, W., Liu W., Huang X-P., E. Vardy, J. McCorvy, X. Gao, X.E. Zhou, K. Melcher, C. Zhang, F.Bai, H. Yang, L. Yang, H. Jiang, B.L. Roth, V. Cherezov, RC. Stevens, and X.E. Xu. Structural Basis for Molecular Recognition at Serotonin Receptors. Science, 2013, 340:610-614.
|
|
